Predicting zoonotic potential of viruses: where are we?
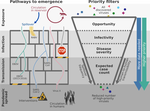
The prospect of identifying high-risk viruses and designing interventions to pre-empt their emergence into human populations is enticing, but controversial, particularly when used to justify large-scale virus discovery initiatives. We review the current state of these efforts, point out unaddressed gaps, and argue that efforts to predict zoonoses provide unique research value that goes beyond their stated – and as yet unrealised – goal of pandemic prevention.
More
Network embedding unveils the hidden interactions in the mammalian virome
We describe a novel bipartite network prediction method, applying it to a global database of mammal-virus interactions. Using predictions from this method to supplement our current limited knowledge of virus host range improves the accuracy of predictions of human infection from viral genome features.
More
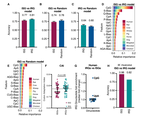
Variation in the ACE2 receptor has limited utility for SARS-CoV-2 host prediction
A large number of publications have shown suprisingly accurate predictions of which species are susceptible to SARS-CoV-2 infection using just the variation in ACE2 proteins found in different species. This would seem to imply that receptor-binding is the primary barrier to cross-species transmission for this virus. However, we show that the predictive power of ACE2-based models derives from strong correlations with host phylogeny rather than processes which can be mechanistically linked to infection biology. Further, biased availability of ACE2 sequences leads to misleading projections of the number and geographic distribution of at-risk species. Models based on host phylogeny reduce this bias, but identify a very large number of susceptible species, implying that model predictions must be combined with local knowledge of exposure risk to practically guide surveillance.
More
The apparent interferon resistance of transmitted HIV-1 is possibly a consequence of enhanced replicative fitness
HIV-1 transmission involves a strong bottleneck, with new infections established by a very small number of virions. These founder viruses appear to be more resistant to interferon than viruses present at later stages of infection, but how (and why) this could be consistently selected for during transmission has remained unclear. We show that, instead of there being a specific molecular mechanism, such apparent resistance may be a simple consequence of how faster-growing viruses behave in typical inteferon-resistance experiments. Bottlenecks are expected to select for faster-growing viruses, but small differences in growth rate may not be apparent in typical growth curve experiments. Adding a growth rate inhibitor such as interferon (but also antiretroviral drugs targeting replication) slows down the entire experiment, allowing the lag phase to be observed and thus emphasising small growth rate differences.
More
Ecological determinants of rabies virus dynamics in vampire bats and spillover to livestock
The epidemiological dynamics of virus transmission among bats remains poorly understood, even for relatively well-studied viruses like rabies. Using longitudinal serology assays of Peruvian vampire bats, we show that rabies maintenance appears to require transmission among multiple, nearby bat colonies which may be facilitated by waning of protective immunity.
More
Bats host the most virulent—but not the most dangerous—zoonotic viruses
Efforts to assess zoonotic risk often use trait-based analyses to identify which viral and reservoir host groups are most likely to source zoonoses, but have not fully addressed how and why the impacts of zoonotic viruses vary in terms of disease severity, capacity to spread within human populations, or total human mortality. We show that, while viruses transmitted by bats are more virulent to humans than average, viruses from close relatives (e.g., primates) are more transmissibile. Importantly, a disproportionately high human death burden is not associated with any animal reservoir, including bats.
More
Mammal virus diversity estimates are unstable due to accelerating discovery effort
While host-virus association data form the backbone of a growing body of comparative work in virus ecology and evolution, knowledge of the wildlife virome is inherently constrained by historical discovery effort. This has led to concern about the reliability of conclusions drawn from currently-available data. By analysing a dataset of discovery dates for 6,571 unique mammal host-virus associations, we show that inference of relative viral richness across host species has been unstable over time. We therefore advise caution to avoid overinterpreting patterns in current data.
More
The science of the host–virus network
Review proposing a framework which unifies a number of recent approaches for understanding and predicting human and animal susceptibility to viral infections.
More
Identifying and prioritizing potential human-infecting viruses from their genome sequences
We describe a machine learning model that identifies candidate zoonoses using evolutionary signals of host range encoded in viral genomes. This allows identification of high-risk viruses immediately upon discovery, increasing both the feasibility and likelihood of downstream virological and ecological characterization and allowing for evidence-driven virus surveillance.
More
The future of zoonotic risk prediction
Findings of an interdisciplinary workshop on zoonotic risk technologies. We discuss the prerequisites, in terms of open data, equity and interdisciplinary collaboration, needed to develop and apply zoonotic risk prediction technologies, the effects such technologies could have (both positive and negative), and how to maximise impacts and ensure equitable access in this rapidly developing field.
More
The antiviral state has shaped the CpG composition of the vertebrate interferome to avoid self-targeting
How do vertebrates evolve sequence-specific antiviral defences without accidentally targeting their own gene transcripts? We show that self-targeting by antiviral effectors has shaped the composition of host transcripts in the vertebrate interferome. These unique compositional signatures give us a better picture of what viral genomes capable of evading sequence-specific host defences might look like, an observation we are exploiting in our work to develop genome-based zoonotic risk prediction methods.
More
Characterizing and evaluating the zoonotic potential of novel viruses discovered in vampire bats
A case study in evaluating the zoonotic potential of newly-discoved viruses using molecular sequencing data. We highlight the value of evaluating zoonotic potential beyond ad hoc consideration of phylogeny and provide surveillance recommendations for novel viruses in a wildlife host which has frequent contact with humans and domestic animals.
More
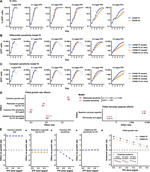
Virulence mismatches in index hosts shape the outcomes of cross-species transmission
Why most cross-species transmissions fail to establish ongoing transmission in the newly infected species remains poorly understood. Examining cross-species inoculations involving rabies, we show that mismatches in virulence which are predictable from host and viral factors make sustained transmission in the novel host less likely. These mechanistic insights help to explain and predict host shift events and highlight meta-analyses of existing experimental inoculation data as a powerful and generalisable approach for understanding the dynamics of index infections in novel species.
More
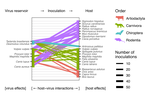
Viral zoonotic risk is homogenous among taxonomic orders of mammalian and avian reservoir hosts
Do some reservoir groups (e.g. bats) produce more zoonotic viruses than others? By cataloguing the accepted reservoirs for 415 viruses associated with mammals and birds, we show that there is currently no evidence for the existence of any such special reservoir groups. Instead, groups containing more reservoir species are associated with more viruses, and proportionally more zoonotic viruses.
More
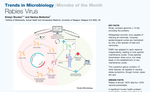
Rabies virus
A mini review for Trends in Microbiology’s “Microbe of the Month” series, describing the transmission cycle of rabies virus in domestic dogs and highlighting the importance of a One Health approach to control and eliminate human rabies deaths.
More
The role of viral evolution in rabies host shifts and emergence
Rabies virus persists in species-specific cycles that rarely sustain transmission in alternative species. The determinants of these species-associations and the adaptive significance of genetic divergence between host-associated virus lineages are poorly understood. This review assesses various lines of evidence and proposes a synthetic hypothesis for the respective roles of ecology and evolution in rabies virus host shifts.
More
A Bayesian approach for inferring the dynamics of partially observed endemic infectious diseases from space-time-genetic data
We describe a statistical framework for reconstructing the sequence of transmission events between observed cases of an endemic infectious disease using genetic, temporal and spatial information. When applied to partial genome sequences of rabies virus sampled from an endemic region of South Africa, our method reveals several distinct transmission cycles with little contact between them, and direct transmission over long distances suggesting significant anthropogenic influence in the movement of infected dogs.
More
Dog rabies in southern Africa: regional surveillance and phylogeographical analyses are an important component of control and elimination strategies
In the resource-poor settings where dog rabies remains endemic, the demonstration of a need to divert scarce funds towards exhaustive …
More