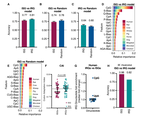
Identifying and prioritizing potential human-infecting viruses from their genome sequences
We describe a machine learning model that identifies candidate zoonoses using evolutionary signals of host range encoded in viral genomes. This allows identification of high-risk viruses immediately upon discovery, increasing both the feasibility and likelihood of downstream virological and ecological characterization and allowing for evidence-driven virus surveillance.
More
The antiviral state has shaped the CpG composition of the vertebrate interferome to avoid self-targeting
How do vertebrates evolve sequence-specific antiviral defences without accidentally targeting their own gene transcripts? We show that self-targeting by antiviral effectors has shaped the composition of host transcripts in the vertebrate interferome. These unique compositional signatures give us a better picture of what viral genomes capable of evading sequence-specific host defences might look like, an observation we are exploiting in our work to develop genome-based zoonotic risk prediction methods.
More